
|
|
ANTIBODY AND ANTIGEN INFORMATION
? »
|
The antibody and antigen information page lists the antigens and corresponding antibodies that have been used in the Human Protein Atlas project, together with the validation results for the antibodies.
In-house generated antibodies have the prefix "HPA". Antibodies provided by external distributors have the prefix "CAB". Links to providers for the antibodies are listed under antibody information, as well as the product name of the antibody as given by the provider.
The host species and the clonality (pAb = polyclonal, mAb = monoclonal, msAb = monospecific) of the antibody,
as well as the purification method is reported. The Human Protein Atlas version in which the antibody was first released is given.
The antigen type, length (amino acid residues), and sequence is reported (if available) under antigen
information. The percent identity (if >=80%) of the antigen sequence to the different proteins encoded by this gene,
and a link to the corresponding protein on the Ensembl
website are listed. Any matches with >=80% sequence identity to proteins from other Ensembl human genes are reported as "Other gene match" together with a link to that gene in the Human Protein Atlas and the maximum sequence identity (%) of the antigen to the matched protein(s) encoded by that gene.
The antibody validation section reports the results from different assays. For immunohistochemistry, the image shown is a selected tissue with representative staining
for this antibody.
This image is clickable for enlarged view. Images from all tissues analysed using this antibody are found by clicking any tissue under the Normal
tissue heading found to the left in the TISSUE subatlas tab, and correspondingly by clicking any
cancer type under the Cancer tissue headings in the menu to the left in the CANCER subatlas tab.
There is also a short summary describing the observed staining patterns using this antibody for immunohistochemistry in a panel of human tissues
(see annotation). The validation scoring of the antibody is described here.
For Western blot, the images for supportive and uncertain validations are shown ( validation description), and the different lanes are described ( WB description). The image is clickable for enlarged view. The expected target mass for the targeted proteins is also listed.
For in-house generated (HPA) antibodies, a protein array assay is performed. The image is a diagram showing the specificity analysis using a number of different Protein Epitope Signature Tags (PrESTs), including the target PrEST (marked in green). The validation is described here.
For the subcellular analysis using immunoflourescence, a representative image from the validation is shown, together with a summary of the staining.
The validation is described here.
|
| |
|
|
Antibody HPA051244
|
|
Antibody CAB002973
|
|
Antibody CAB039238
|
|
Antibody CAB039239
|
|
Antibody CAB072876
|
|
|
ANTIBODY INFORMATION
|
|
Provider |
Atlas Antibodies
Sigma-Aldrich
| | DakoCytomation
| | Lab Vision/NeoMarkers
| | Lab Vision/NeoMarkers
| | Atlas Antibodies
Sigma-Aldrich
| |
Product name |
HPA051244 | | M7001 | | MS-187 | | MS-104 | | AMAb90956 | |
Host species |
Rabbit | | Mouse | | Mouse | | Mouse | | Mouse | |
Clonality |
pAb | | mAb | | mAb | | mAb | | mAb | |
Purity |
Affinity purified using the PrEST-antigen as affinity ligand | | Not known | | Protein A/G | | Protein A/G | | Protein A/G | |
Released in version |
14 | | 1 | | 7 | | 7 | | 14 | |
|
ANTIGEN INFORMATION
|
|
Antigen |
Recombinant protein fragment | | Recombinant protein | | Recombinant protein | | Recombinant protein | | Recombinant protein | |
Length (aa) |
89 | | | | | | | | | |
Antigen sequence |
KKGEPHHELPPGSTKRALPNNTSSSPQPKKKPLDGEYFTLQIRGRERFEM
FRELNEALELKDAQAGKEPGGSRAHSSHLKSKKGQSTSR
| |
| |
| |
| |
| |
Matching transcripts |
TP53-001 - ENSP00000269305 [100%]
TP53-002 - ENSP00000391478 [100%]
TP53-008 - ENSP00000481179 [100%]
TP53-021 - ENSP00000481638 [100%]
TP53-022 - ENSP00000478219 [100%]
TP53-023 - ENSP00000484375 [100%]
TP53-028 - ENSP00000482537 [100%]
TP53-201 - ENSP00000482903 [100%]
| | | | | | | | | |
Other gene match |
| | | | | | | | | |
|
ANTIBODY VALIDATION
|
|
|
Immunohistochemistry
|
|
Image |
| |  | |  | |  | |  | |
Description |
Application not done for this antibody. | | Immunohistochemical staining of human skin shows nuclear positivity in subsets of cells.
More information | | Immunohistochemical staining of human skin shows nuclear positivity.
More information | | Immunohistochemical staining of human skin shows nuclear positivity in subsets of cells.
More information | | Immunohistochemical staining of human colon shows no positivity in glandular cells.
More information | |
Retrieval method |
| | HIER pH6 | | HIER pH6 | | HIER pH6 | | HIER pH6 | |
Antibody dilution |
| | 1:5000 | | 1:4000 | | 1:250 | | 1:200 | |
Literature conformity |
| | Consistent with extensive gene/protein characterization data | | Consistent with extensive gene/protein characterization data | | Consistent with extensive gene/protein characterization data | | Consistent with extensive gene/protein characterization data | |
RNA consistency |
| | Mainly consistent with RNA expression data | | Mainly consistent with RNA expression data | | Mainly consistent with RNA expression data | | Mainly consistent with RNA expression data | |
|
Immunofluorescence
|
|
Image |
 | | | |  | | 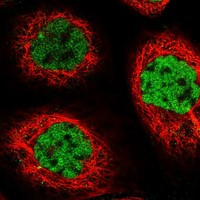 | | | |
Description |
Immunofluorescent staining of human cell line A-431 shows positivity in nucleus but excluded from the nucleoli.
More information | | Application not done for this antibody. | | Immunofluorescent staining of human cell line A-431 shows positivity in nucleus but excluded from the nucleoli.
More information | | Immunofluorescent staining of human cell line A-431 shows positivity in nucleus but excluded from the nucleoli.
More information | | Application not done for this antibody. | |
Antibody dilution |
1:78 | | | | 1:150 | | 1:75 | | | |
Validation IF |
Supportive: The subcellular location is supported by experimental gene/protein characterization data, gene silencing, or an independent antibody. | | | | Supportive: The subcellular location is supported by experimental gene/protein characterization data, gene silencing, or an independent antibody. | | Supportive: The subcellular location is supported by experimental gene/protein characterization data, gene silencing, or an independent antibody. | | | |
|
siRNA
|
|
Image |
 | | | | | | | | | |
Description |
Signal downregulation > 25% by one siRNA.
More information | | Application not done for this antibody. | | Application not done for this antibody. | | Application not done for this antibody. | | Application not done for this antibody. | |
Antibody dilution |
1:78 | | | | | | | | | |
Validation siRNA |
Supportive: The siRNA validation is supportive. | | | | | | | | | |
|
Western Blot
|
|
Image |
 | |  | |  | | 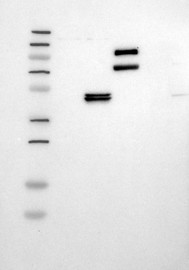 | |  | |
Description |
Lane 1: Marker [kDa] 250, 130, 95, 72, 55, 36, 28, 17, 10
Lane 2: RT4
Lane 3: U-251 MG
Lane 4: Human Plasma
Lane 5: Liver
Lane 6: Tonsil
More information | | Lane 1: Marker [kDa] 229, 107, 78, 46, 31, 26, 16.2
Lane 2: RT4
Lane 3: U-251 MG
Lane 4: Human Plasma
Lane 5: Liver
Lane 6: Tonsil
More information | | Lane 1: Marker [kDa] 250, 130, 95, 72, 55, 36, 28, 17, 11
Lane 2: RT4
Lane 3: U-251 MG
Lane 4: Human Plasma
Lane 5: Liver
Lane 6: Tonsil
More information | | Lane 1: Marker [kDa] 250, 130, 95, 72, 55, 36, 28, 17, 11
Lane 2: RT4
Lane 3: U-251 MG
Lane 4: Human Plasma
Lane 5: Liver
Lane 6: Tonsil
More information | | Lane 1: Marker [kDa] 250, 130, 95, 72, 55, 36, 28, 17, 10
Lane 2: RT4
Lane 3: U-251 MG
Lane 4: Human Plasma
Lane 5: Liver
Lane 6: Tonsil
More information | |
Target mass (kDa) |
43.7, 42.6, 39.3, 29.6, 26.6 | | 43.7, 42.6, 39.3, 38.5, 37.9, 37.8, 34.2, 33.5, 31.5, 29.6, 26.6, 24.4, 23.7, 22.5, 21.4, 20.7, 17.8, 17.7, 15.4, 13.7, 3.4 | | 43.7, 42.6, 39.3, 38.5, 37.9, 37.8, 34.2, 33.5, 31.5, 29.6, 26.6, 24.4, 23.7, 22.5, 21.4, 20.7, 17.8, 17.7, 15.4, 13.7, 3.4 | | 43.7, 42.6, 39.3, 38.5, 37.9, 37.8, 34.2, 33.5, 31.5, 29.6, 26.6, 24.4, 23.7, 22.5, 21.4, 20.7, 17.8, 17.7, 15.4, 13.7, 3.4 | | 43.7, 42.6, 39.3, 38.5, 37.9, 37.8, 34.2, 33.5, 31.5, 29.6, 26.6, 24.4, 23.7, 22.5, 21.4, 20.7, 17.8, 17.7, 15.4, 13.7, 3.4 | |
Antibody dilution |
1:390 | | 1:500 | | 1:250 | | 1:250 | | 1:500 | |
Validation WB |
Supportive: Single band corresponding to the predicted size in kDa (+/-20%) | | Supportive: Band of predicted size in kDa (+/-20%) with additional bands present | | Supportive: Band of predicted size in kDa (+/-20%) with additional bands present | | Supportive: Band of predicted size in kDa (+/-20%) with additional bands present | | Uncertain: Single band differing more than +/-20% from predicted size in kDa and not supported by experimental and/or bioinformatic data | |
|
Protein array
|
|
Image |
 | | | | | | | | | |
Description |
Antibody specificity analysis with protein arrays. Predicted and matching interactions are shown in green.
More information | | Application not done for this antibody. | | Application not done for this antibody. | | Application not done for this antibody. | | Application not done for this antibody. | |
Antibody dilution |
1:3100 | | | | | | | | | |
Validation PA |
Supportive: Pass with single peak corresponding to interaction only with its own antigen. | | | | | | | | | |
| |
|
|
ANTIGEN VIEW
? »
|
The antigen view displays the antigen location on the target protein(s) and the features of the target protein. The tabs at the top of the protein view section can be used to switch between the different splice variants to which an antigen has been mapped.
At the top of the antigen view, the position of the antigen (identified by the corresponding HPA identifier) is shown as a green bar. A yellow triangle on the bar indicates a <100% sequence identity to the protein target.
Under the antigens, the maximum percent sequence identity of the protein to all other proteins from other human genes is displayed, using a sliding window of 10 aa residues (HsID 10) or 50 aa residues (HsID 50) ( read more).
If a signal peptide is predicted by a majority of the signal peptide predictors
SPOCTOPUS,
SignalP 4.0, and
Phobius (turquoise) and/or transmembrane regions (orange) are predicted by MDM, these are displayed.
Low complexity regions are shown in yellow and InterPro regions in green.
Common (purple) and unique (grey) regions between alternative processed transcripts from the same gene are also displayed ( read more), and at the bottom of the protein view is the protein scale.
|
|
|
|
|
|
|
|